## Ages
Xmx <- 70
Xmn <- 15
##Generate Sample Data
dat_sim <- function(n=1000){
X <- seq(Xmn, Xmx, length.out=n)
## Random Productivity
e <- runif(n, 0, 1E6)
beta <- 1E-10*exp(1.4*X -.015*X^2)
Y <- (beta*X + e)/10
return(data.frame(Y, X))
}
dat0 <- dat_sim(1000)
dat0 <- dat0[order(dat0[, 'X']), ]
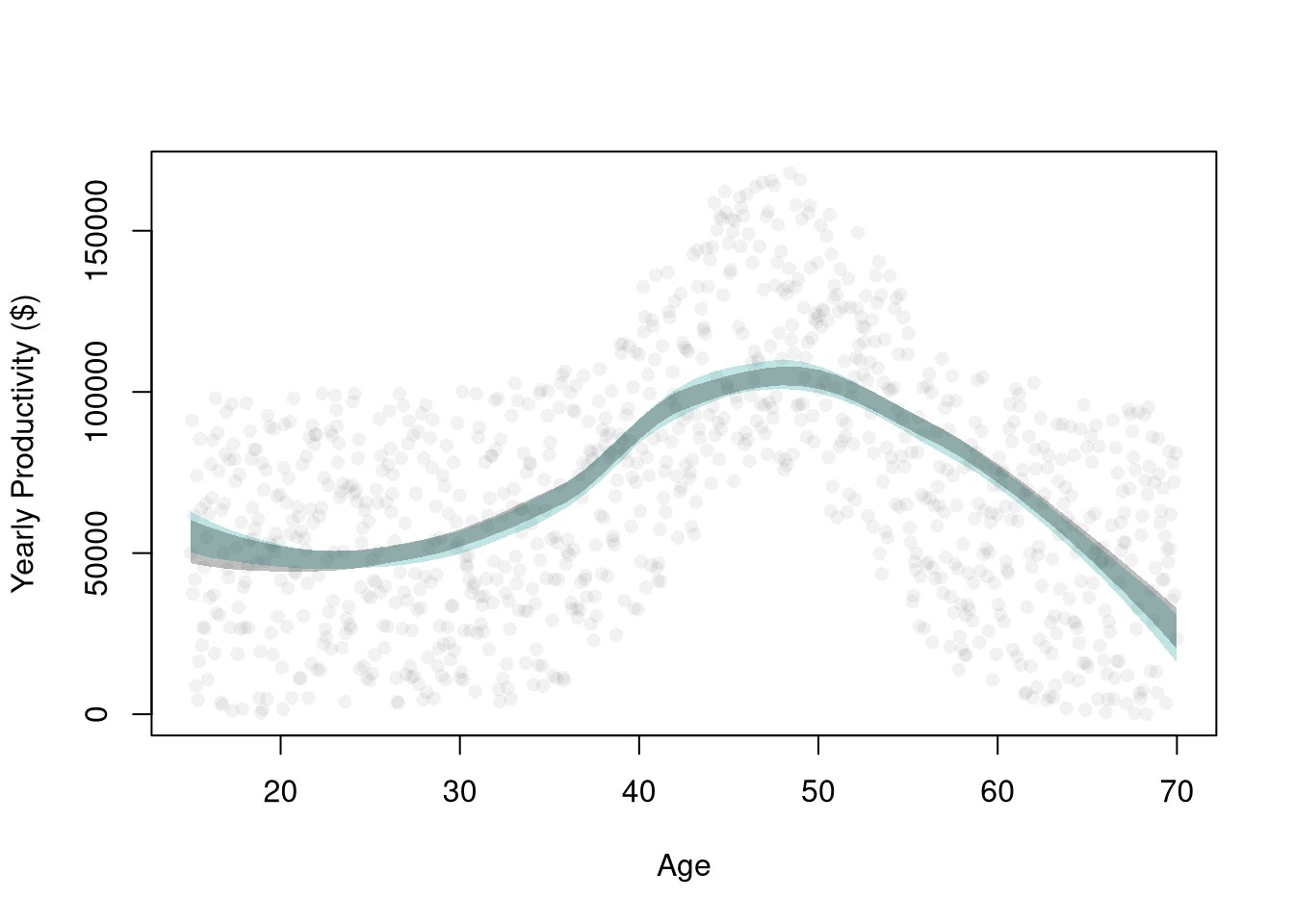
## Data from one sample
plot(Y ~ X, data=dat0, pch=16, col=grey(0, .05),
main=NA, ylab='Yearly Productivity ($)', xlab='Age' )
reg_lo <- loess(Y ~ X, data=dat0, span=.8)
## Plot Bootstrap CI for Single Sample
pred_design <- data.frame(X=seq(Xmn, Xmx))
preds_lo <- predict(reg_lo, newdata=pred_design)
boot_lo <- matrix(NA, nrow=nrow(pred_design), ncol=399)
for (b in 1:399) {
dat0_i <- dat0[sample(nrow(dat0), replace=T), ]
reg_i <- loess(Y ~ X, data=dat0_i, span=.8)
boot_lo[, b] <- predict(reg_i, newdata=pred_design)
}
boot_cb <- apply(boot_lo, 1, quantile,
probs=c(.025, .975), na.rm=T)
polygon(
c(pred_design[[1]], rev(pred_design[[1]])),
c(boot_cb[1, ], rev(boot_cb[2, ])),
col=hcl.colors(3, alpha=.25)[2],
border=NA)
# Construct CI across Multiple Samples
sample_lo <- matrix(NA, nrow=nrow(pred_design), ncol=399)
for (b in 1:399) {
xy_b <- dat_sim(1000) #Entirely new sample
reg_b <- loess(Y ~ X, data=xy_b, span=.8)
sample_lo[, b] <- predict(reg_b, newdata=pred_design)
}
ci_lo <- apply(sample_lo, 1, quantile,
probs=c(.025, .975), na.rm=T)
polygon(
c(pred_design[[1]], rev(pred_design[[1]])),
c(ci_lo[1, ], rev(ci_lo[2, ])),
col=grey(0, alpha=.25),
border=NA)