library(mvtnorm)
# Simulate Bivariate Data
N <- 10000
Mu <- c(2, 2) ## Means
Sigma1 <- matrix(c(2, -.8, -.8, 1), 2, 2) ## CoVariance Matrix 1
MVdat1 <- rmvnorm(N, Mu, Sigma1)
colnames(MVdat1) <- c('X', 'Y')
Sigma2 <- matrix(c(2, .4, .4, 1), 2, 2) ## CoVariance Matrix 2
MVdat2 <- rmvnorm(N, Mu, Sigma2)
colnames(MVdat2) <- c('X', 'Y')
par(mfrow=c(1, 2))
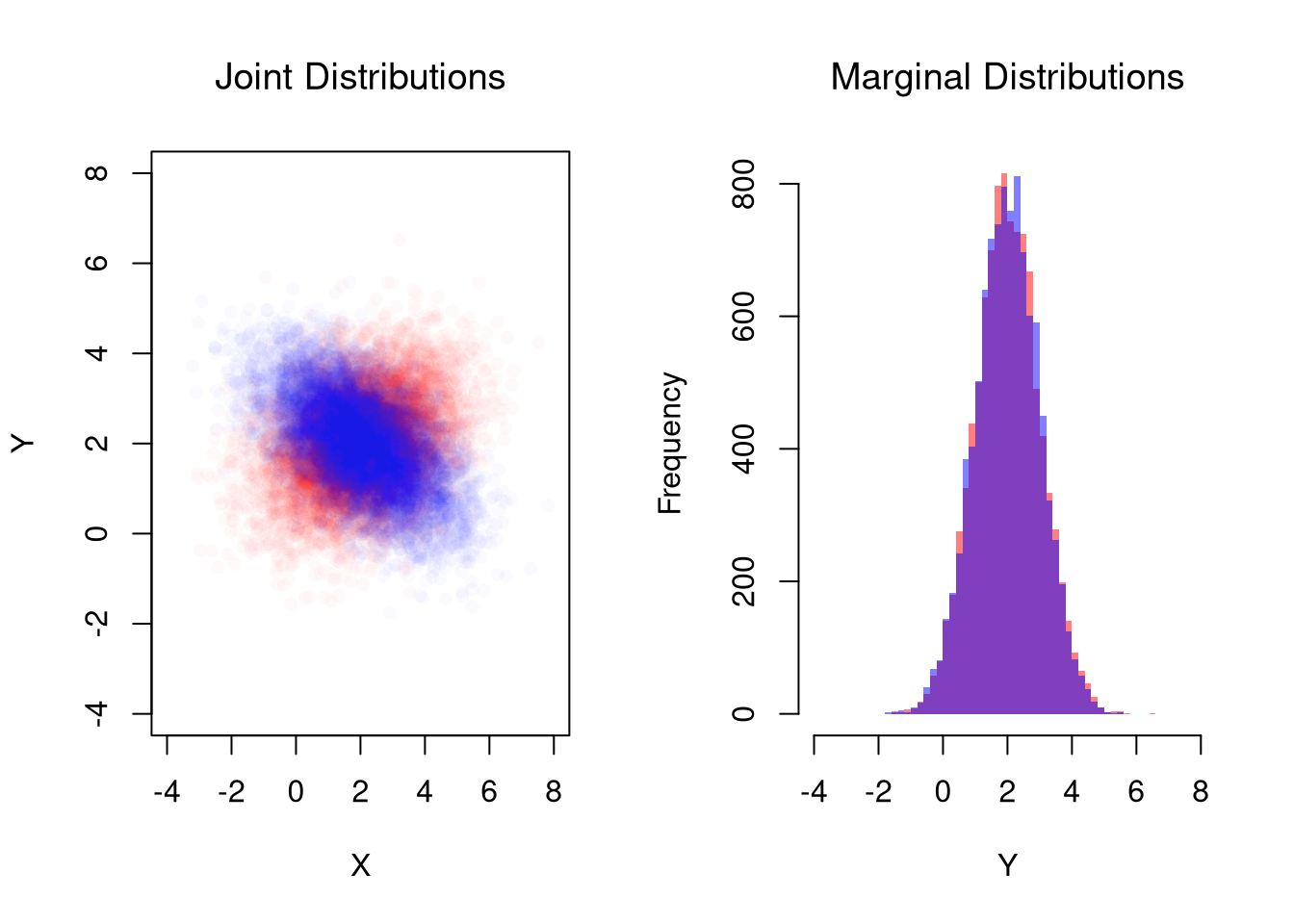
## Different diagonals
plot(MVdat2, col=rgb(1, 0, 0, 0.02), pch=16,
main=NA,
ylim=c(-4, 8), xlim=c(-4, 8),
xlab='X', ylab='Y')
title('Joint Distributions', font.main=1)
points(MVdat1, col=rgb(0, 0, 1, 0.02), pch=16)
## Same marginal distributions
xbks <- seq(-4, 8, by=.2)
hist(MVdat2[, 2], col=rgb(1, 0, 0, 0.5),
breaks=xbks, border=NA, freq=F,
xlab='Y',
main=NA)
title('Marginal Distributions', font.main=1)
hist(MVdat1[, 2], col=rgb(0, 0, 1, 0.5),
add=T, breaks=xbks, border=NA, freq=F)